Rendering seamless 3D panoramas
Exploring Blender’s panoramic camera modes to make a seamless circular render.
Hi! I’m Christopher, a seventh-year PhD student in the Galloway Lab. You can find my publications and posts below. Check out the About page for contact information, Github, ORCID, and more.
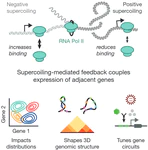
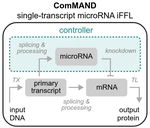

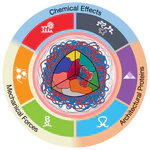
Exploring Blender’s panoramic camera modes to make a seamless circular render.
